From Surf Wiki (app.surf) — the open knowledge base
Ribosomal DNA
Genes coding for ribosomal RNA
Genes coding for ribosomal RNA
The ribosomal DNA (rDNA) consists of a group of ribosomal RNA encoding genes and related regulatory elements, and is widespread in similar configuration in all domains of life. The ribosomal DNA encodes the non-coding ribosomal RNA, integral structural elements in the assembly of ribosomes, its importance making it the most abundant section of RNA found in cells of eukaryotes. Additionally, these segments include regulatory sections, such as a promoter specific to the RNA polymerase I, as well as both transcribed and non-transcribed spacer segments.
Due to their high importance in the assembly of ribosomes for protein biosynthesis, the rDNA genes are usually highly conserved in molecular evolution. The number of copies can vary considerably per species. Ribosomal DNA is widely used for phylogenetic studies.
Structure
| Type | SSU rRNA | LSU rRNA |
|---|---|---|
| Eukaryotes | 18S rRNA | 28S rRNA |
| 5.8S rRNA | ||
| 5S rRNA | ||
| Bacteria | 16S rRNA | 23S rRNA |
| 5S rRNA | ||
| Mitochondrial | MT-RNR1 (12S rRNA) | MT-RNR2 (16S rRNA) |
| Plastid | 16S rRNA | 23S rRNA |
| 4.5S rRNA | ||
| 5S rRNA |
The ribosomal DNA includes all genes coding for the non-coding structural ribosomal RNA molecules. Across all domains of life, these are the structural sequences of the small subunit (16S or 18S rRNA) and the large subunit (23S or 28S rRNA). The assembly of the latter also include the 5S rRNA as well as the additional 5.8S rRNA in eukaryotes.
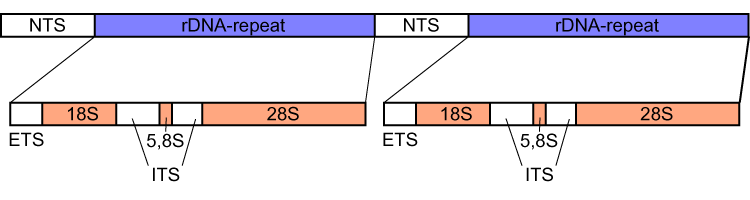
The rDNA-genes are commonly present with multiple copies in the genome, where they are organized in linked groups in most species, separated by an internal transcribed spacer (ITS) and preceded by the external transcribed spacer (ETS). The 5S rRNA is also linked to these rDNA region in prokaryotes, while it is located in separate repeating regions in most eukaryotes. They are transcribed together to a precursor RNA which is then processed to equal amounts of each rRNA.
Prokaryotes
The primary structural rRNA molecules in Bacteria and Archaea are smaller than their counterparts in eukaryotes, grouped as 16S rRNA and 23S rRNA. Meanwhile, the 5S rRNA also present in prokaryotes, is of a similar size to eukaryotes.
The form of rDNA operon most bacteria and archaea is "linked": from 3' to 5' one sees a continuous tract of 16S–23S–5S. The part between 16S and 23S is called the internal transcribed spacer (ITS) and often includes a tRNA. The part between 23S and 5S, though technically also a spacer that is internal and transcribed, does not have a name of its own.
A notable amount of bacteria and archaea diverge from the canonical structure of the operon containing the rDNA genes, instead carrying them as "unlinked" 16S and 23S–5S genes in different places of their genome. Archaea also show some other forms of divergence: the Thermoproteati have an eukaryote-style operon without 5S, and some archaeons have all three segments in different sites.
Plastids
Ribosomal DNA in typical chloroplasts follows the canonical structure of prokaryotic ribosomal DNA and goes: 16S–trnI–trnA–23S–4.5S–5S. The 4.5S corresponds to the 5' end fragment of 23S in bacteria.
The human microchondrial DNA shows a typical 12S–tRNAVal–16S organization, where 12S and 16S are greatly reduced versions of 16S and 23S of bacteria. Most vertebrates have the same organization of the rDNA operon, as do ticks. Some eukaryotes such as snails have a split structure where 16S and 12S are separate.
Eukaryotes

NTS, nontranscribed spacer, ETS, external transcribed spacer, ITS, internal transcribed spacers 1 and 2, numbered from 5' end.]]
.png)
The 45S rDNA gene cluster of eukaryotes consists of the genes for the 18S, 5.8S and 28S rRNA, separated by the two ITS-1 and ITS-2 spacers. The active genome of eukaryotes contains several hundred copies of the polycistronic rDNA transcriptional unit as tandem repeats, they are organized in nucleolus organizer regions (NORs), which in turn can be present at multiple loci in the genome.
Similar to the structure of prokaryotes, the 5S rRNA is appended to the rDNA cluster in the Saccharomycetes 5S rDNA is also present in independent tandem repeats as in Drosophila.
As repetitive DNA regions often undergo recombination events, the rDNA repeats have many regulatory mechanisms that keep the DNA from undergoing mutations, thus keeping the rDNA conserved.
In the nucleus, the nucleolus organizer regions give rise to the nucleolus, where the rDNA regions of the chromosome forms expanded chromosomal loops, accessible for transcription of rRNA. In rDNA, the tandem repeats are mostly found in the nucleolus; but heterochromatic rDNA is found outside of the nucleolus. However, transcriptionally active rDNA resides inside of the nucleolus itself.
Humans
The human genome contains a total of 560 copies
Human 5S ribosomal DNA is located on chromosome 1. There are 17 copies in the human reference genome, in loci coded RNA5S1 through RNA5S17. An average haploid human genome actually has 98 copies of 5S.
Ciliates
In ciliates, the presence of a generative micronucleus next to the vegetative macronucleus allows for the reduction of rDNA genes in the germline. The exact number of copies in the micronucleus core genome ranging from several copies in Paramecium as low as a single copy in Tetrahymena thermophila Similar processes are reported from Glaucoma chattoni and to lesser extent from Paramecium.
Sequence homogeneity
In the large rDNA array, polymorphisms between rDNA repeat units are very low, indicating that rDNA tandem arrays are evolving through concerted evolution.
5S tandem repeat sequences in several Drosophila were compared with each other; the result revealed that insertions and deletions occurred frequently between species and often flanked by conserved sequences. They could occur by slippage of the newly synthesized strand during DNA replication or by gene conversion.
Sequence divergence
The rDNA transcription tracts have low rate of polymorphism among species, which allows interspecific comparison to elucidate phylogenetic relationship using only a few specimens. Coding regions of rDNA are highly conserved among species but ITS regions are variable due to insertions, deletions, and point mutations. Between remote species as human and frog comparison of sequences at ITS tracts is not appropriate. Conserved sequences at coding regions of rDNA allow comparisons of remote species, even between yeast and human. Human 5.8S rRNA has 75% identity with yeast 5.8S rRNA. In cases for sibling species, comparison of the rDNA segment including ITS tracts among species and phylogenetic analysis are made satisfactorily. The different coding regions of the rDNA repeats usually show distinct evolutionary rates. As a result, this DNA can provide phylogenetic information of species belonging to wide systematic levels.
Recombination-stimulating activity in yeast
A fragment of yeast rDNA containing the 5S gene, non-transcribed spacer DNA, and part of the 25S (yeast version of 28S) gene has localized cis-acting mitotic recombination stimulating activity. This DNA fragment contains a mitotic recombination hotspot, referred to as HOT1. HOT1 expresses recombination-stimulating activity when it is inserted into novel locations in the yeast genome. HOT1 includes an RNA polymerase I (PolI) transcription promoter that catalyzes 35S ribosomal rRNA (yeast version of 45S) gene transcription. In a PolI defective mutant, the HOT1 hotspot recombination-stimulating activity is abolished. The level of PolI transcription in HOT1 appears to determine the level of recombination.
Clinical significance
Diseases can be associated with DNA mutations where DNA can be expanded, such as Huntington's disease, or lost due to deletion mutations. The same is true for mutations that occur in rDNA repeats; it has been found that if the genes that are associated with the synthesis of ribosomes are disrupted or mutated, it can result in various diseases associated with the skeleton or bone marrow. Also, any damage or disruption to the enzymes that protect the tandem repeats of the rDNA, can result in lower synthesis of ribosomes, which also lead to other defects in the cell. Neurological diseases can also arise from mutations in the rDNA tandem repeats, such as Bloom syndrome, which occurs when the number of tandem repeats increases close to a hundred-fold; compared with that of the normal number of tandem repeats. Various types of cancers can also be born from mutations of the tandem repeats in the ribosomal DNA. Cell lines can become malignant from either a rearrangement of the tandem repeats, or an expansion of the repeats in the rDNA.
References
References
- (December 1991). "Ribosomal DNA: Molecular Evolution and Phylogenetic Inference". The Quarterly Review of Biology.
- (January 1991). "16S ribosomal DNA amplification for phylogenetic study". Journal of Bacteriology.
- Graw, Jochen. (2015). "Genetik". Springer-Verlag Berlin Heidelberg.
- (22 July 2021). "Ribosome Biogenesis in Archaea". Frontiers in Microbiology.
- (February 2020). "Unlinked rRNA genes are widespread among bacteria and archaea". The ISME Journal.
- (14 May 2024). "Chloroplast Ribosome Biogenesis Factors". Plant and Cell Physiology.
- (April 1981). "Sequence and organization of the human mitochondrial genome". Nature.
- (2 January 2016). "The complete mitochondrial genome of the salmon shark, Lamna ditropis (Chondrichthyes, Lamnidae)". Mitochondrial DNA Part A.
- (11 November 2022). "Amplification and sequencing of entire tick mitochondrial genomes for a phylogenomic analysis". Scientific Reports.
- (2013). "The complete mitochondrial genome of the land snail Cornu aspersum (Helicidae: Mollusca): intra-specific divergence of protein-coding genes and phylogenetic considerations within Euthyneura.". PLOS ONE.
- (2 July 2024). "Mitochondrial genome of the fluke pond snail, Austropeplea cf. brazieri (Gastropoda: Lymnaeidae)". Parasites & Vectors.
- (9 May 2018). "Complete mitochondrial and rDNA complex sequences of important vector species of Biomphalaria, obligatory hosts of the human-infecting blood fluke, Schistosoma mansoni.". Scientific Reports.
- (December 2008). "Comparative genomics and molecular dynamics of DNA repeats in eukaryotes". Microbiology and Molecular Biology Reviews.
- (2019-03-01). "Keeping ribosomal DNA intact: a repeating challenge". Chromosome Research.
- Schmidt, Olaf. (2017). "Genetik und Molekularbiologie". Springer-Verlag Berlin Heidelberg.
- "Gene group {{!}} HUGO Gene Nomenclature Committee".
- (January 2008). "Genomic architecture and inheritance of human ribosomal RNA gene clusters.". Genome Research.
- (2015). "Intragenomic polymorphisms among high-copy loci: a genus-wide study of nuclear ribosomal DNA in Asclepias (Apocynaceae)". PeerJ.
- (November 1995). "Structural evolution of the Drosophila 5S ribosomal genes". Journal of Molecular Evolution.
- (April 2004). "Sequencing and analysis of the internal transcribed spacers (ITSs) and coding regions in the EcoR I fragment of the ribosomal DNA of the Japanese pond frog Rana nigromaculata". Genes & Genetic Systems.
- (February 1976). "Sequence homologies in mammalian 5.8S ribosomal RNA". Biochemistry.
- (1998). "Sequence differences of rDNA-ITS2 and species-diagnostic PCR assay of ''Anopheles sinensis'' and ''Anopheles anthropophagus'' from China". J Med Coll PLA.
- (2005). "A newly recognized species in the ''Anopheles'' hyrcanus group and molecular identification of related species from the Republic of South Korea (Diptera: Culicidae)". Zootaxa.
- (December 1984). "Cis-acting, recombination-stimulating activity in a fragment of the ribosomal DNA of S. cerevisiae". Cell.
- (April 2004). "Transcription-mediated hyper-recombination in HOT1". Genes to Cells.
- (March 2019). "Keeping ribosomal DNA intact: a repeating challenge". Chromosome Research.
This article was imported from Wikipedia and is available under the Creative Commons Attribution-ShareAlike 4.0 License. Content has been adapted to SurfDoc format. Original contributors can be found on the article history page.
Ask Mako anything about Ribosomal DNA — get instant answers, deeper analysis, and related topics.
Research with MakoFree with your Surf account
Create a free account to save articles, ask Mako questions, and organize your research.
Sign up freeThis content may have been generated or modified by AI. CloudSurf Software LLC is not responsible for the accuracy, completeness, or reliability of AI-generated content. Always verify important information from primary sources.
Report