From Surf Wiki (app.surf) — the open knowledge base
Nicotinamide adenine dinucleotide
Chemical compound which is reduced and oxidized
Chemical compound which is reduced and oxidized
C21H29N7O14P2 (reduced) 665.4 g/mol (reduced) | NFPA-H = 1 | NFPA-F = 1 | NFPA-R = 0
Nicotinamide adenine dinucleotide (NAD) is a coenzyme central to metabolism. Found in all living cells, NAD is called a dinucleotide because it consists of two nucleotides joined through their phosphate groups. One nucleotide contains an adenine nucleobase and the other, nicotinamide. NAD exists in two forms: an oxidized and reduced form, abbreviated as NAD and NADH (H for hydrogen), respectively.
In cellular metabolism, NAD is involved in redox reactions, carrying electrons from one reaction to another, so it is found in two forms: NAD is an oxidizing agent, accepting electrons from other molecules and becoming reduced; with H+, this reaction forms NADH, which can be used as a reducing agent to donate electrons. These electron transfer reactions are the main function of NAD. It is also used in other cellular processes, most notably as a substrate of enzymes in adding or removing chemical groups to or from proteins, in posttranslational modifications. Because of the importance of these functions, the enzymes involved in NAD metabolism are targets for drug discovery.
In organisms, NAD can be synthesized from simple building-blocks (de novo) from either tryptophan or aspartic acid, each a case of an amino acid. Alternatively, more complex components of the coenzymes are taken up from nutritive compounds such as vitamin B3 (also called niacin, or (nicotinic acid, hence the name "nicotinamide"). Similar compounds are produced by reactions that break down the structure of NAD, providing a salvage pathway that recycles them back into their respective active form.
In the name NAD, the superscripted plus sign indicates the positive formal charge on one of its nitrogen atoms.
Some NAD is converted into the coenzyme nicotinamide adenine dinucleotide phosphate (NADP), whose chemistry largely parallels that of NAD, though its predominant role is as a coenzyme in anabolic metabolism. NADP is a reducing agent in anabolic reactions like the Calvin cycle and lipid and nucleic acid syntheses. NADP exists in two forms: NADP+, the oxidized form, and NADPH, the reduced form. NADP is similar to nicotinamide adenine dinucleotide (NAD), but NADP has a phosphate group at the C-2′ position of the adenosyl.
Physical and chemical properties
Nicotinamide adenine dinucleotide consists of two nucleosides joined by pyrophosphate. The nucleosides each contain a ribose ring, one with adenine attached to the first carbon atom (the 1' position) (adenosine diphosphate ribose) and the other with nicotinamide at this position.
The compound accepts or donates the equivalent of H−. Such reactions (summarized in formula below) involve the removal of two hydrogen atoms from a reactant (R), in the form of a hydride ion (H−), and a proton (H). The proton is released into solution, while the reductant RH is oxidized and NAD reduced to NADH by transfer of the hydride to the nicotinamide ring.
:RH + NAD → NADH + H + R;
From the electron pair of the hydride ion, one electron is attracted to the slightly more electronegative atom of the nicotinamide ring of NAD, becoming part of the nicotinamide moiety. The remaining hydrogen atom is transferred to the carbon atom opposite the N atom. The midpoint potential of the NAD/NADH redox pair is −0.32 volts, which makes NADH a moderately strong reducing agent. The reaction is easily reversible, when NADH reduces another molecule and is re-oxidized to NAD. This means the coenzyme can continuously cycle between the NAD and NADH forms without being consumed.
In appearance, all forms of this coenzyme are white amorphous powders that are hygroscopic and highly water-soluble. The solids are stable if stored dry and in the dark. Solutions of NAD are colorless and stable for about a week at 4 °C and neutral pH, but decompose rapidly in acidic or alkaline solutions. Upon decomposition, they form products that are enzyme inhibitors.
Both NAD and NADH strongly absorb ultraviolet light because of the adenine. For example, peak absorption of NAD is at a wavelength of 259 nanometers (nm), with an extinction coefficient of 16,900 M−1cm−1. NADH also absorbs at higher wavelengths, with a second peak in UV absorption at 339 nm with an extinction coefficient of 6,220 M−1cm−1. This difference in the ultraviolet absorption spectra between the oxidized and reduced forms of the coenzymes at higher wavelengths makes it simple to measure the conversion of one to another in enzyme assays – by measuring the amount of UV absorption at 340 nm using a spectrophotometer.
NAD and NADH also differ in their fluorescence. Freely diffusing NADH in aqueous solution, when excited at the nicotinamide absorbance of ~335 nm (near-UV), fluoresces at 445–460 nm (violet to blue) with a fluorescence lifetime of 0.4 nanoseconds, while NAD does not fluoresce. The properties of the fluorescence signal changes when NADH binds to proteins, so these changes can be used to measure dissociation constants, which are useful in the study of enzyme kinetics. These changes in fluorescence are also used to measure changes in the redox state of living cells, through fluorescence microscopy.
NADH can be converted to NAD+ in a reaction catalysed by copper, which requires hydrogen peroxide. Thus, the supply of NAD+ in cells requires mitochondrial copper(II).
Concentration and state in cells
In rat liver, the total amount of NAD and NADH is approximately 1 μmole per gram of wet weight, about 10 times the concentration of NADP and NADPH in the same cells. The actual concentration of NAD in cell cytosol is harder to measure, with recent estimates in animal cells ranging around 0.3 mM, and approximately 1.0 to 2.0 mM in yeast. However, more than 80% of NADH fluorescence in mitochondria is from bound form, so the concentration in solution is much lower.
NAD concentrations are highest in the mitochondria, constituting 40% to 70% of the total cellular NAD. NAD in the cytosol is carried into the mitochondrion by a specific membrane transport protein, since the coenzyme cannot diffuse across membranes. The intracellular half-life of NAD+ was claimed to be between 1–2 hours by one review, whereas another review gave varying estimates based on compartment: intracellular 1–4 hours, cytoplasmic 2 hours, and mitochondrial 4–6 hours.
The balance between the oxidized and reduced forms of nicotinamide adenine dinucleotide is called the NAD/NADH ratio. This ratio is an important component of what is called the redox state of a cell, a measurement that reflects both the metabolic activities and the health of cells. The effects of the NAD/NADH ratio are complex, controlling the activity of several key enzymes, including glyceraldehyde 3-phosphate dehydrogenase and pyruvate dehydrogenase. In healthy mammalian tissues, estimates of the ratio of free NAD to NADH in the cytoplasm typically lie around 700:1; the ratio is thus favorable for oxidative reactions. The ratio of total NAD/NADH is much lower, with estimates ranging from 3–10 in mammals. In contrast, the NADP/NADPH ratio is normally about 0.005, so NADPH is the dominant form of this coenzyme. These different ratios are key to the different metabolic roles of NADH and NADPH.
Biosynthesis
NAD is synthesized through two metabolic pathways. It is produced either in a de novo pathway from amino acids or in salvage pathways by recycling preformed components such as nicotinamide back to NAD. Although most tissues synthesize NAD by the salvage pathway in mammals, much more de novo synthesis occurs in the liver from tryptophan, and in the kidney and macrophages from nicotinic acid.
''De novo'' production
Most organisms synthesize NAD from simple components. The quinolinic acid is converted to nicotinic acid mononucleotide (NaMN) by transfer of a phosphoribose moiety. An adenylate moiety is then transferred to form nicotinic acid adenine dinucleotide (NaAD). Finally, the nicotinic acid moiety in NaAD is amidated to a nicotinamide (Nam) moiety, forming nicotinamide adenine dinucleotide.
In a further step, some NAD is converted into NADP by NAD kinase, which phosphorylates NAD. In most organisms, this enzyme uses adenosine triphosphate (ATP) as the source of the phosphate group, although several bacteria such as Mycobacterium tuberculosis and a hyperthermophilic archaeon Pyrococcus horikoshii, use inorganic polyphosphate as an alternative phosphoryl donor.
Salvage pathways
Despite the presence of the de novo pathway, the salvage reactions are essential in humans; a lack of vitamin B3 in the diet causes the vitamin deficiency disease pellagra. This high requirement for NAD results from the constant consumption of the coenzyme in reactions such as posttranslational modifications, since the cycling of NAD between oxidized and reduced forms in redox reactions does not change the overall levels of the coenzyme. The major source of NAD in mammals is the salvage pathway which recycles the nicotinamide produced by enzymes utilizing NAD. The first step, and the rate-limiting enzyme in the salvage pathway is nicotinamide phosphoribosyltransferase (NAMPT), which produces nicotinamide mononucleotide (NMN). NMN is the immediate precursor to NAD+ in the salvage pathway.
Besides assembling NAD de novo from simple amino acid precursors, cells also salvage preformed compounds containing a pyridine base. The three vitamin precursors used in these salvage metabolic pathways are nicotinic acid (NA), nicotinamide (Nam) and nicotinamide riboside (NR). There are some reports that mammalian cells can take up extracellular NAD from their surroundings, and both nicotinamide and nicotinamide riboside can be absorbed from the gut.
The salvage pathways used in microorganisms differ from those of mammals. Some pathogens, such as the yeast Candida glabrata and the bacterium Haemophilus influenzae are NAD auxotrophs – they cannot synthesize NAD – but possess salvage pathways and thus are dependent on external sources of NAD or its precursors. Even more surprising is the intracellular pathogen Chlamydia trachomatis, which lacks recognizable candidates for any genes involved in the biosynthesis or salvage of both NAD and NADP, and must acquire these coenzymes from its host.
Functions
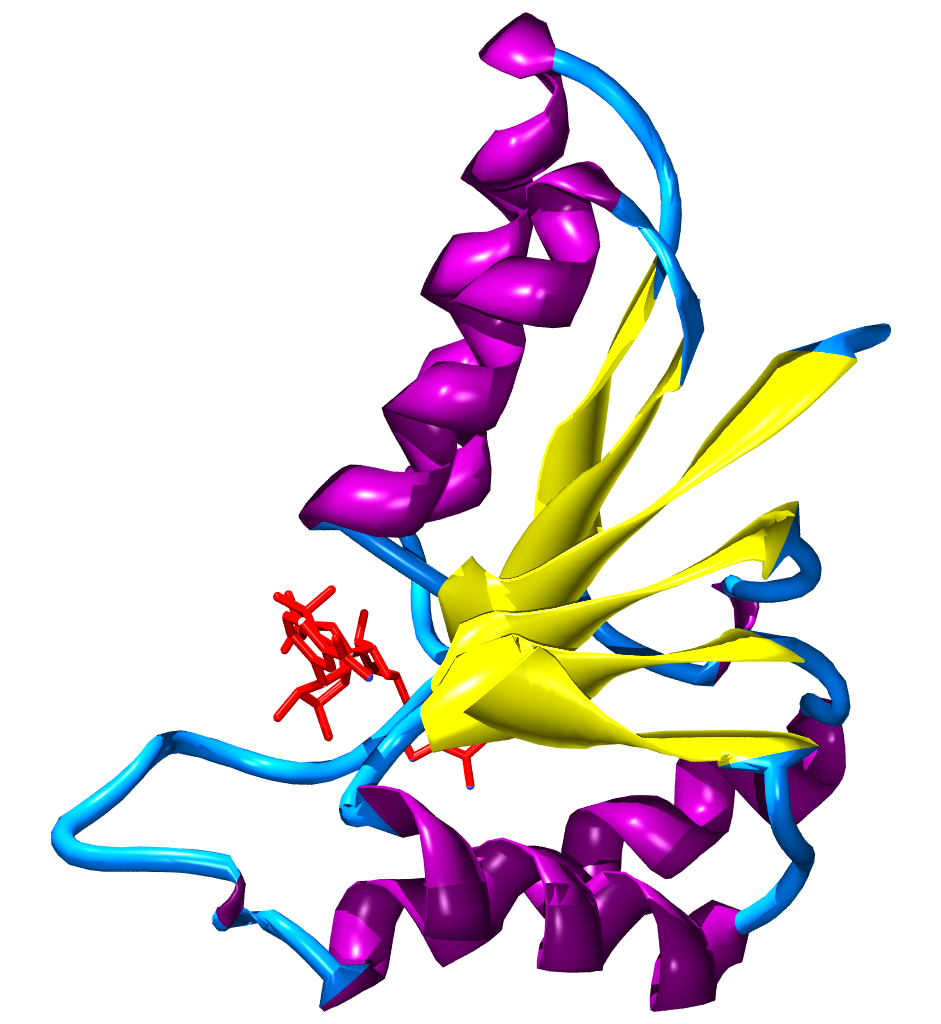
Nicotinamide adenine dinucleotide has several essential roles in metabolism. It acts as a coenzyme in redox reactions, as a donor of ADP-ribose moieties in ADP-ribosylation reactions, as a precursor of the second messenger molecule cyclic ADP-ribose, as well as acting as a substrate for bacterial DNA ligases and a group of enzymes called sirtuins that use NAD to remove acetyl groups from proteins. In addition to these metabolic functions, NAD+ emerges as an adenine nucleotide that can be released from cells spontaneously and by regulated mechanisms, and can therefore have important extracellular roles.
Oxidoreductase binding of NAD
The main role of NAD in metabolism is the transfer of electrons from one molecule to another. Reactions of this type are catalyzed by a large group of enzymes called oxidoreductases. The correct names for these enzymes contain the names of both their substrates: for example NADH-ubiquinone oxidoreductase catalyzes the oxidation of NADH by coenzyme Q. However, these enzymes are also referred to as dehydrogenases or reductases, with NADH-ubiquinone oxidoreductase commonly being called NADH dehydrogenase or sometimes coenzyme Q reductase.
There are many different superfamilies of enzymes that bind NAD / NADH. One of the most common superfamilies includes a structural motif known as the Rossmann fold. The motif is named after Michael Rossmann, who was the first scientist to notice how common this structure is within nucleotide-binding proteins.
An example of a NAD-binding bacterial enzyme involved in amino acid metabolism that does not have the Rossmann fold is found in Pseudomonas syringae pv. tomato (; ).
When bound in the active site of an oxidoreductase, the nicotinamide ring of the coenzyme is positioned so that it can accept a hydride from the other substrate. Depending on the enzyme, the hydride donor is positioned either "above" or "below" the plane of the planar C4 carbon, as defined in the figure. Class A oxidoreductases transfer the atom from above; class B enzymes transfer it from below. Since the C4 carbon that accepts the hydrogen is prochiral, this can be exploited in enzyme kinetics to give information about the enzyme's mechanism. This is done by mixing an enzyme with a substrate that has deuterium atoms substituted for the hydrogens, so the enzyme will reduce NAD by transferring deuterium rather than hydrogen. In this case, an enzyme can produce one of two stereoisomers of NADH.
Despite the similarity in how proteins bind the two coenzymes, enzymes almost always show a high level of specificity for either NAD or NADP. This specificity reflects the distinct metabolic roles of the respective coenzymes, and is the result of distinct sets of amino acid residues in the two types of coenzyme-binding pocket. For instance, in the active site of NADP-dependent enzymes, an ionic bond is formed between a basic amino acid side-chain and the acidic phosphate group of NADP. On the converse, in NAD-dependent enzymes the charge in this pocket is reversed, preventing NADP from binding. However, there are a few exceptions to this general rule, and enzymes such as aldose reductase, glucose-6-phosphate dehydrogenase, and methylenetetrahydrofolate reductase can use both coenzymes in some species.
Role in redox metabolism
The redox reactions catalyzed by oxidoreductases are vital in all parts of metabolism, but one particularly important area where these reactions occur is in the release of energy from nutrients. Here, reduced compounds such as glucose and fatty acids are oxidized, thereby releasing energy. This energy is transferred to NAD by reduction to NADH, as part of beta oxidation, glycolysis, and the citric acid cycle. In eukaryotes the electrons carried by the NADH that is produced in the cytoplasm are transferred into the mitochondrion (to reduce mitochondrial NAD) by mitochondrial shuttles, such as the malate-aspartate shuttle. The mitochondrial NADH is then oxidized in turn by the electron transport chain, which pumps protons across a membrane and generates ATP through oxidative phosphorylation. These shuttle systems also have the same transport function in chloroplasts.
Since both the oxidized and reduced forms of nicotinamide adenine dinucleotide are used in these linked sets of reactions, the cell maintains significant concentrations of both NAD and NADH, with the high NAD/NADH ratio allowing this coenzyme to act as both an oxidizing and a reducing agent. In contrast, the main function of NADPH is as a reducing agent in anabolism, with this coenzyme being involved in pathways such as fatty acid synthesis and photosynthesis. Since NADPH is needed to drive redox reactions as a strong reducing agent, the NADP/NADPH ratio is kept very low.
Although it is important in catabolism, NADH is also used in anabolic reactions, such as gluconeogenesis. This need for NADH in anabolism poses a problem for prokaryotes growing on nutrients that release only a small amount of energy. For example, nitrifying bacteria such as Nitrobacter oxidize nitrite to nitrate, which releases sufficient energy to pump protons and generate ATP, but not enough to produce NADH directly. As NADH is still needed for anabolic reactions, these bacteria use a nitrite oxidoreductase to produce enough proton-motive force to run part of the electron transport chain in reverse, generating NADH.
Non-redox roles
The coenzyme NAD is also consumed in ADP-ribose transfer reactions. For example, enzymes called ADP-ribosyltransferases add the ADP-ribose moiety of this molecule to proteins, in a posttranslational modification called ADP-ribosylation. ADP-ribosylation involves either the addition of a single ADP-ribose moiety, in mono-ADP-ribosylation, or the transferral of ADP-ribose to proteins in long branched chains, which is called poly(ADP-ribosyl)ation. Mono-ADP-ribosylation was first identified as the mechanism of a group of bacterial toxins, notably cholera toxin, but it is also involved in normal cell signaling. Poly(ADP-ribosyl)ation is carried out by the poly(ADP-ribose) polymerases. The poly(ADP-ribose) structure is involved in the regulation of several cellular events and is most important in the cell nucleus, in processes such as DNA repair and telomere maintenance. NAD may also be added onto cellular RNA as a 5'-terminal modification.
Another function of this coenzyme in cell signaling is as a precursor of cyclic ADP-ribose, which is produced from NAD by ADP-ribosyl cyclases, as part of a second messenger system. This molecule acts in calcium signaling by releasing calcium from intracellular stores. It does this by binding to and opening a class of calcium channels called ryanodine receptors, which are located in the membranes of organelles, such as the endoplasmic reticulum, and inducing the activation of the transcription factor NAFC3
NAD is also consumed by different NAD+-consuming enzymes, such as CD38, CD157, PARPs and the NAD-dependent deacetylases (sirtuins, such as Sir2.). These enzymes act by transferring an acetyl group from their substrate protein to the ADP-ribose moiety of NAD; this cleaves the coenzyme and releases nicotinamide and O-acetyl-ADP-ribose. The sirtuins mainly seem to be involved in regulating transcription through deacetylating histones and altering nucleosome structure. However, non-histone proteins can be deacetylated by sirtuins as well. These activities of sirtuins are particularly interesting because of their importance in the regulation of aging.
Other NAD-dependent enzymes include bacterial DNA ligases, which join two DNA ends by using NAD as a substrate to donate an adenosine monophosphate (AMP) moiety to the 5' phosphate of one DNA end. This intermediate is then attacked by the 3' hydroxyl group of the other DNA end, forming a new phosphodiester bond. This contrasts with eukaryotic DNA ligases, which use ATP to form the DNA-AMP intermediate.
Li et al. have found that NAD directly regulates protein-protein interactions. They also show that one of the causes of age-related decline in DNA repair may be increased binding of the protein DBC1 (Deleted in Breast Cancer 1) to PARP1 (poly[ADP–ribose] polymerase 1) as NAD levels decline during aging. The decline in cellular concentrations of NAD during aging likely contributes to the aging process and to the pathogenesis of the chronic diseases of aging. Thus, the modulation of NAD may protect against cancer, radiation, and aging.
Extracellular actions of NAD+
In recent years, NAD+ has also been recognized as an extracellular signaling molecule involved in cell-to-cell communication. NAD+ is released from neurons in blood vessels, large intestine, from neurosecretory cells, and from brain synaptosomes, and is proposed to be a novel neurotransmitter that transmits information from nerves to effector cells in smooth muscle organs. Further studies are needed to determine the underlying mechanisms of its extracellular actions and their importance for human health and life processes in other organisms.
Clinical significance
The enzymes that make and use NAD and NADH are important in both pharmacology and the research into future treatments for disease. Drug design and drug development exploits NAD in three ways: as a direct target of drugs, by designing enzyme inhibitors or activators based on its structure that change the activity of NAD-dependent enzymes, and by trying to inhibit NAD biosynthesis.
Because cancer cells utilize increased glycolysis, and because NAD enhances glycolysis, nicotinamide phosphoribosyltransferase (NAD salvage pathway) is often amplified in cancer cells.
It has been studied for its potential use in the therapy of neurodegenerative diseases such as Alzheimer's and Parkinson's disease as well as multiple sclerosis. A placebo-controlled clinical trial of NADH (which excluded NADH precursors) in people with Parkinson's failed to show any effect.
NAD is also a direct target of the drug isoniazid, which is used in the treatment of tuberculosis, an infection caused by Mycobacterium tuberculosis. Isoniazid is a prodrug and once it has entered the bacteria, it is activated by a peroxidase enzyme, which oxidizes the compound into a free radical form. This radical then reacts with NADH, to produce adducts that are very potent inhibitors of the enzymes enoyl-acyl carrier protein reductase, and dihydrofolate reductase.
Since many oxidoreductases use NAD and NADH as substrates, and bind them using a highly conserved structural motif, the idea that inhibitors based on NAD could be specific to one enzyme is surprising. However, this can be possible: for example, inhibitors based on the compounds mycophenolic acid and tiazofurin inhibit IMP dehydrogenase at the NAD binding site. Because of the importance of this enzyme in purine metabolism, these compounds may be useful as anti-cancer, anti-viral, or immunosuppressive drugs. Other drugs are not enzyme inhibitors, but instead activate enzymes involved in NAD metabolism. Sirtuins are a particularly interesting target for such drugs, since activation of these NAD-dependent deacetylases extends lifespan in some animal models. Compounds such as resveratrol increase the activity of these enzymes, which may be important in their ability to delay aging in both vertebrate, and invertebrate model organisms. In one experiment, mice given NAD for one week had improved nuclear-mitochrondrial communication.
Because of the differences in the metabolic pathways of NAD biosynthesis between organisms, such as between bacteria and humans, this area of metabolism is a promising area for the development of new antibiotics. For example, the enzyme nicotinamidase, which converts nicotinamide to nicotinic acid, is a target for drug design because this enzyme is absent in humans but present in yeast and bacteria.
In bacteriology, NAD, sometimes referred to factor V, is used as a supplement to culture media for some fastidious bacteria.
High-cost unlicensed infusions of NAD+ have been claimed in the UK to be "clinically proven" and "effective" treatment for alcoholism and drug abuse. NAD+ is not approved or licensed for medical use in the UK; there are likely breaches of advertising and medicines rules, and no proof that treatments work. Medical experts say "It's complete nonsense" ... "It's untested and unproven. We don't know anything about its efficacy or long-term safety". A November 2024 study, cited 700 times, claiming that NAD+ levels in lab rats decreased with age was withdrawn after images were found to have been manipulated, and underlying data was not provided to the publishers on request.
History
The coenzyme NAD was first discovered by the British biochemists Arthur Harden and William John Young in 1906. They noticed that adding boiled and filtered yeast extract greatly accelerated alcoholic fermentation in unboiled yeast extracts. They called the unidentified factor responsible for this effect a coferment. Through a long and difficult purification from yeast extracts, this heat-stable factor was identified as a nucleotide sugar phosphate by Hans von Euler-Chelpin. In 1936, the German scientist Otto Heinrich Warburg showed the function of the nucleotide coenzyme in hydride transfer and identified the nicotinamide portion as the site of redox reactions.
Vitamin precursors of NAD were first identified in 1938, when Conrad Elvehjem showed that liver has an "anti-black tongue" activity in the form of nicotinamide. Then, in 1939, he provided the first strong evidence that nicotinic acid is used to synthesize NAD. In the early 1940s, Arthur Kornberg was the first to detect an enzyme in the biosynthetic pathway. In 1949, the American biochemists Morris Friedkin and Albert L. Lehninger proved that NADH linked metabolic pathways such as the citric acid cycle with the synthesis of ATP in oxidative phosphorylation. In 1958, Jack Preiss and Philip Handler discovered the intermediates and enzymes involved in the biosynthesis of NAD; salvage synthesis from nicotinic acid is termed the Preiss-Handler pathway. In 2004, Charles Brenner and co-workers uncovered the nicotinamide riboside kinase pathway to NAD.
The non-redox roles of NAD(P) were discovered later. Studies in the 1980s and 1990s revealed the activities of NAD and NADP metabolites in cell signaling – such as the action of cyclic ADP-ribose, which was discovered in 1987.
The metabolism of NAD remained an area of intense research into the 21st century, with interest heightened after the discovery of the NAD-dependent protein deacetylases called sirtuins in 2000, by Shin-ichiro Imai and coworkers in the laboratory of Leonard P. Guarente. In 2009 Imai proposed the "NAD World" hypothesis that key regulators of aging and longevity in mammals are sirtuin 1 and the primary NAD synthesizing enzyme nicotinamide phosphoribosyltransferase (NAMPT). In 2016 Imai expanded his hypothesis to "NAD World 2.0", which postulates that extracellular NAMPT from adipose tissue maintains NAD in the hypothalamus (the control center) in conjunction with myokines from skeletal muscle cells. In 2018, Napa Therapeutics was formed to develop drugs against a novel aging-related target based on the research in NAD metabolism conducted in the lab of Eric Verdin.
References
References
- {{Lehninger4th
- The nicotinamide group can be attached in two orientations to the anomeric ribose carbon atom. Because of these two possible structures, the NAD could exists as either of two [[diastereomer]]s. It is the β-nicotinamide diastereomer of NAD{{+ that is found in nature.
- (2007). "The power to reduce: pyridine nucleotides – small molecules with a multitude of functions". Biochem. J..
- (January 2007). "NAD+ metabolism in health and disease". Trends in Biochemical Sciences.
- (25 June 2025). "What is NAD?".
- (1997). "Alternative respiratory pathways of ''Escherichia coli'': energetics and transcriptional regulation in response to electron acceptors". Biochim. Biophys. Acta.
- Windholz, Martha. (1983). "The Merck Index: an encyclopedia of chemicals, drugs, and biologicals". Merck.
- (1979). "Structure of lactate dehydrogenase inhibitor generated from coenzyme". Biochemistry.
- Dawson, R. Ben. (1985). "Data for biochemical research". Clarendon Press.
- (2014-05-29). "Separating NADH and NADPH fluorescence in live cells and tissues using FLIM". Springer Science and Business Media LLC.
- (1992). "Fluorescence lifetime imaging of free and protein-bound NADH". Proc. Natl. Acad. Sci. U.S.A..
- (1989). "Time-resolved fluorescence studies on NADH bound to mitochondrial malate dehydrogenase". Biochim. Biophys. Acta.
- (2006). "The Free NADH Concentration Is Kept Constant in Plant Mitochondria under Different Metabolic Conditions". Plant Cell.
- (September 1980). "Copper (II) complex-catalyzed oxidation of NADH by hydrogen peroxide". Biol Trace Elem Res.
- (2023). "A druggable copper-signalling pathway that drives inflammation". Nature.
- (1984). "Measurement of tissue purine, pyrimidine, and other nucleotides by radial compression high-performance liquid chromatography". Anal. Biochem..
- (2006). "The simultaneous measurement of nicotinamide adenine dinucleotide and related compounds by liquid chromatography/electrospray ionization tandem mass spectrometry". Anal. Biochem..
- (2007). "Nutrient-Sensitive Mitochondrial NAD+ Levels Dictate Cell Survival". Cell.
- (2007). "Nicotinamide riboside promotes Sir2 silencing and extends lifespan via Nrk and Urh1/Pnp1/Meu1 pathways to NAD+". Cell.
- (2005). "Distribution of mitochondrial NADH fluorescence lifetimes: steady-state kinetics of matrix NADH interactions". Biochemistry.
- (2019). "Regulation of Glucose Metabolism by NAD + and ADP-Ribosylation". [[Cells (journal).
- (2006). "Identification of the mitochondrial NAD+ transporter in ''Saccharomyces cerevisiae''". J. Biol. Chem..
- Srivastava S. (2016). "Emerging therapeutic roles for NAD(+) metabolism in mitochondrial and age-related disorders". Clinical and Translational Medicine.
- (2018). "Sirtuins in Health and Disease".
- (2001). "Redox environment of the cell as viewed through the redox state of the glutathione disulfide/glutathione couple". Free Radic Biol Med.
- (1967). "The redox state of free nicotinamide-adenine dinucleotide in the cytoplasm and mitochondria of rat liver". Biochem. J..
- (2002). "Regulation of corepressor function by nuclear NADH". Science.
- (April 2003). "Nicotinamide adenine dinucleotide, a metabolic regulator of transcription, longevity and disease". Curr. Opin. Cell Biol..
- (1969). "The redox state of free nicotinamide–adenine dinucleotide phosphate in the cytoplasm of rat liver". Biochem. J..
- (2020). "Age-related NAD + decline". [[Experimental Gerontology]].
- (2006). "Early Steps in the Biosynthesis of NAD in Arabidopsis Start with Aspartate and Occur in the Plastid". Plant Physiol..
- (1 March 1980). "Nicotinamide adenine dinucleotide biosynthesis and pyridine nucleotide cycle metabolism in microbial systems". Microbiol. Rev..
- (2006). "Structural and functional properties of NAD kinase, a key enzyme in NADP biosynthesis". Mini Reviews in Medicinal Chemistry.
- (2005). "First Archaeal Inorganic Polyphosphate/ATP-Dependent NAD Kinase, from Hyperthermophilic Archaeon Pyrococcus horikoshii: Cloning, Expression, and Characterization". Appl. Environ. Microbiol..
- (2004). "Characterization of Mycobacterium tuberculosis NAD kinase: functional analysis of the full-length enzyme by site-directed mutagenesis". Biochemistry.
- Henderson LM. (1983). "Niacin". Annu. Rev. Nutr..
- (2018). "Therapeutic Potential of NAD-Boosting Molecules: The In Vivo Evidence". [[Cell Metabolism]].
- "What is NMN?".
- (2008). "Characterization of NAD Uptake in Mammalian Cells". J. Biol. Chem..
- (2016). "Nicotinamide riboside is uniquely and orally bioavailable in mice and humans". Nature Communications.
- (2003). "Reconstructing eukaryotic NAD metabolism". BioEssays.
- (2007). "Assimilation of NAD+ precursors in ''Candida glabrata''". Mol. Microbiol..
- (2000). "NADP and NAD utilization in ''Haemophilus influenzae''". Mol. Microbiol..
- (2002). "From Genetic Footprinting to Antimicrobial Drug Targets: Examples in Cofactor Biosynthetic Pathways". J. Bacteriol..
- (2005). "Crystallization of three key glycolytic enzymes of the opportunistic pathogen ''Cryptosporidium parvum''". Biochim. Biophys. Acta.
- (2004). "Release of beta-nicotinamide adenine dinucleotide upon stimulation of postganglionic nerve terminals in blood vessels and urinary bladder". J Biol Chem.
- (2006). "Emerging functions of extracellular pyridine nucleotides". Mol. Med..
- "Enzyme Nomenclature, Recommendations for enzyme names from the Nomenclature Committee of the International Union of Biochemistry and Molecular Biology".
- "NiceZyme View of ENZYME: EC 1.6.5.3". Expasy.
- (2015). "Proteopedia: Rossmann fold: A beta-alpha-beta fold at dinucleotide binding sites". Biochem Mol Biol Educ.
- Lesk AM. (1995). "NAD-binding domains of dehydrogenases". Curr. Opin. Struct. Biol..
- (1973). "Comparison of super-secondary structures in proteins". J Mol Biol.
- (2005). "Crystal structures of Delta1-piperideine-2-carboxylate/Delta1-pyrroline-2-carboxylate reductase belonging to a new family of NAD(P)H-dependent oxidoreductases: conformational change, substrate recognition, and stereochemistry of the reaction". J. Biol. Chem..
- Bellamacina CR. (1 September 1996). "The nicotinamide dinucleotide binding motif: a comparison of nucleotide binding proteins". FASEB J..
- (1997). "NADP-dependent enzymes. I: Conserved stereochemistry of cofactor binding". Proteins.
- (2006). "Biochemical and genetic analysis of methylenetetrahydrofolate reductase in Leishmania metabolism and virulence". J. Biol. Chem..
- (2001). "Stoichiometry and compartmentation of NADH metabolism in ''Saccharomyces cerevisiae''". FEMS Microbiol. Rev..
- (1 December 2003). "The molecular machinery of Keilin's respiratory chain". Biochemical Society Transactions.
- (1991). "Redox Transfer across the Inner Chloroplast Envelope Membrane". Plant Physiol.
- Nicholls DG. (2002). "Bioenergetics 3". Academic Press.
- (October 1985). "The interaction between the cytosolic pyridine nucleotide redox potential and gluconeogenesis from lactate/pyruvate in isolated rat hepatocytes. Implications for investigations of hormone action.". Journal of Biological Chemistry.
- (1990). "Energy conservation in ''Nitrobacter''". FEMS Microbiology Letters.
- (2006). "Genome Sequence of the Chemolithoautotrophic Nitrite-Oxidizing Bacterium ''Nitrobacter winogradskyi'' Nb-255". Appl. Environ. Microbiol..
- Ziegler M. (2000). "New functions of a long-known molecule. Emerging roles of NAD in cellular signaling". Eur. J. Biochem..
- (2005). "Introduction to poly(ADP-ribose) metabolism". Cell. Mol. Life Sci..
- (2004). "The new life of a centenarian: signaling functions of NAD(P)". Trends Biochem. Sci..
- (2003). "New Embo Member's Review: Functional aspects of protein mono-ADP-ribosylation". EMBO J..
- Bürkle A. (2005). "Poly(ADP-ribose). The most elaborate metabolite of NAD+". FEBS J..
- (2004). "Ecto-ADP-ribosyltransferases (ARTs): emerging actors in cell communication and signaling". Curr. Med. Chem..
- (December 2009). "LC/MS analysis of cellular RNA reveals NAD-linked RNA". Nat Chem Biol.
- Guse AH. (2004). "Biochemistry, biology, and pharmacology of cyclic adenosine diphosphoribose (cADPR)". Curr. Med. Chem..
- Guse AH. (2004). "Regulation of calcium signaling by the second messenger cyclic adenosine diphosphoribose (cADPR)". Curr. Mol. Med..
- Guse AH. (2005). "Second messenger function and the structure-activity relationship of cyclic adenosine diphosphoribose (cADPR)". FEBS J..
- (2004). "Sirtuins: Sir2-related NAD-dependent protein deacetylases". Genome Biol.
- Verdin, Eric. (2015-12-04). "NAD⁺ in aging, metabolism, and neurodegeneration". Science.
- (June 2004). "The Sir2 Family of Protein Deacetylases". Annual Review of Biochemistry.
- (2006). "The role of NAD+ dependent histone deacetylases (sirtuins) in ageing". Curr Drug Targets.
- (2022-08-30). "NAD+ metabolism drives astrocyte proinflammatory reprogramming in central nervous system autoimmunity". Proceedings of the National Academy of Sciences of the United States of America.
- (2001). "Bacterial DNA ligases". Mol. Microbiol..
- (1997). "A newly identified DNA ligase of ''Saccharomyces cerevisiae'' involved in RAD52-independent repair of DNA double-strand breaks". Genes & Development.
- (23 March 2017). "A conserved NAD binding pocket that regulates protein-protein interactions during aging". Science.
- Verdin E. NAD⁺ in aging, metabolism, and neurodegeneration. Science. 2015 Dec 4;350(6265):1208-13. doi: 10.1126/science.aac4854. PMID 26785480
- (2004). "NAD+ surfaces again". Biochem. J..
- (2011). "Compartmentation of NAD+-dependent signalling". FEBS Lett..
- (February 2006). "β-NAD is a novel nucleotide released on stimulation of nerve terminals in human urinary bladder detrusor muscle". American Journal of Physiology. Renal Physiology.
- (2007). "Beta-nicotinamide adenine dinucleotide is an inhibitory neurotransmitter in visceral smooth muscle". Proc. Natl. Acad. Sci. U.S.A..
- (2011). "β-nicotinamide adenine dinucleotide is an enteric inhibitory neurotransmitter in human and nonhuman primate colons". Gastroenterology.
- (2009). "Storage and secretion of beta-NAD, ATP and dopamine in NGF-differentiated rat pheochromocytoma PC12 cells". Eur. J. Neurosci..
- (2012). "Release, neuronal effects and removal of extracellular β-nicotinamide adenine dinucleotide (β-NAD+) in the rat brain". Eur. J. Neurosci..
- (2017). "A lectin receptor kinase as a potential sensor for extracellular nicotinamide adenine dinucleotide in Arabidopsis thaliana". eLife.
- Sauve AA. (March 2008). "NAD+ and vitamin B3: from metabolism to therapies". The Journal of Pharmacology and Experimental Therapeutics.
- (2007). "Nicotinamide adenine dinucleotide metabolism as an attractive target for drug discovery". Expert Opin. Ther. Targets.
- (2018). "NAD Metabolism in Cancer Therapeutics". [[Frontiers in Microbiology]].
- (2020). "NAD- and NADPH-Contributing Enzymes as Therapeutic Targets in Cancer: An Overview". [[Biomolecules (journal).
- Swerdlow RH. (1998). "Is NADH effective in the treatment of Parkinson's disease?". Drugs Aging.
- (2006). "Mechanisms of action of isoniazid". Mol. Microbiol..
- (2003). "The isoniazid-NAD adduct is a slow, tight-binding inhibitor of InhA, the Mycobacterium tuberculosis enoyl reductase: Adduct affinity and drug resistance". Proc. Natl. Acad. Sci. U.S.A..
- (2006). "Mycobacterium tuberculosis dihydrofolate reductase is a target for isoniazid". Nat. Struct. Mol. Biol..
- (2004). "Cofactor mimics as selective inhibitors of NAD-dependent inosine monophosphate dehydrogenase (IMPDH){{snd}}the major therapeutic target". Curr. Med. Chem..
- (1999). "Nucleoside and non-nucleoside IMP dehydrogenase inhibitors as antitumor and antiviral agents". Curr. Med. Chem..
- (2008). "SIRT1: roles in aging and cancer". BMB Rep.
- (2006). "Resveratrol prolongs lifespan and retards the onset of age-related markers in a short-lived vertebrate". Curr. Biol..
- (2003). "Small molecule activators of sirtuins extend ''Saccharomyces cerevisiae'' lifespan". Nature.
- (2004). "Sirtuin activators mimic caloric restriction and delay ageing in metazoans". Nature.
- (19 December 2013). "Declining NAD+ Induces a Pseudohypoxic State Disrupting Nuclear-Mitochondrial Communication during Aging". Cell.
- (2002). "Structural biology of enzymes involved in NAD and molybdenum cofactor biosynthesis". Curr. Opin. Struct. Biol..
- (2001). "Cofactor Biosynthesis".
- (30 March 2021). "Meningitis Lab Manual: ID and Characterization of Hib". CDC.
- Das, Shanti. (23 February 2025). "'It's not ethical and it's not medical': how UK rehab clinics are cashing in on NAD+". The Observer.
- (24 October 1906). "The alcoholic ferment of yeast-juice Part II.--The coferment of yeast-juice". Proceedings of the Royal Society of London.
- "Fermentation of sugars and fermentative enzymes". Nobel Foundation.
- (1936). "Pyridin, der wasserstoffübertragende bestandteil von gärungsfermenten (pyridin-nucleotide)". Biochemische Zeitschrift.
- (1938). "The isolation and identification of the anti-black tongue factor". J. Biol. Chem..
- (1939). "The effect of a nicotinic acid deficiency upon the coenzyme I content of animal tissues". J. Biol. Chem..
- Kornberg A. (1948). "The participation of inorganic pyrophosphate in the reversible enzymatic synthesis of diphosphopyridine nucleotide". J. Biol. Chem..
- (1 April 1949). "Esterification of inorganic phosphate coupled to electron transport between dihydrodiphosphopyridine nucleotide and oxygen". J. Biol. Chem..
- (1958). "Biosynthesis of diphosphopyridine nucleotide. I. Identification of intermediates". J. Biol. Chem..
- (1958). "Biosynthesis of diphosphopyridine nucleotide. II. Enzymatic aspects". J. Biol. Chem..
- Bieganowski, P. (2004). "Discoveries of Nicotinamide Riboside as a Nutrient and Conserved NRK Genes Establish a Preiss-Handler Independent Route to NAD+ in Fungi and Humans". Cell.
- (1963). "Nicotinamide mononucleotide activation of new DNA-dependent polyadenylic acid synthesizing nuclear enzyme". Biochem. Biophys. Res. Commun..
- (15 July 1987). "Pyridine nucleotide metabolites stimulate calcium release from sea urchin egg microsomes desensitized to inositol trisphosphate". J. Biol. Chem..
- (2000). "Transcriptional silencing and longevity protein Sir2 is an NAD-dependent histone deacetylase". Nature.
- Imai S. (2009). "The NAD World: a new systemic regulatory network for metabolism and aging--Sirt1, systemic NAD biosynthesis, and their importance". [[Cell Biochemistry and Biophysics]].
- Imai S. (2016). "The NAD World 2.0: the importance of the inter-tissue communication mediated by NAMPT/NAD +/SIRT1 in mammalian aging and longevity control". [[List of Nature Research journals#N.
- (2018-08-17). "Napa Therapeutics Formed to Develop Drugs to Influence NAD Metabolism".
This article was imported from Wikipedia and is available under the Creative Commons Attribution-ShareAlike 4.0 License. Content has been adapted to SurfDoc format. Original contributors can be found on the article history page.
Ask Mako anything about Nicotinamide adenine dinucleotide — get instant answers, deeper analysis, and related topics.
Research with MakoFree with your Surf account
Create a free account to save articles, ask Mako questions, and organize your research.
Sign up freeThis content may have been generated or modified by AI. CloudSurf Software LLC is not responsible for the accuracy, completeness, or reliability of AI-generated content. Always verify important information from primary sources.
Report