From Surf Wiki (app.surf) — the open knowledge base
Fluorinase

| Field | Value |
|---|---|
| Name | Fluorinase (adenosyl-fluoride synthase) |
| EC_number | 2.5.1.63 |
| image | fluorinase crystal structure.png |
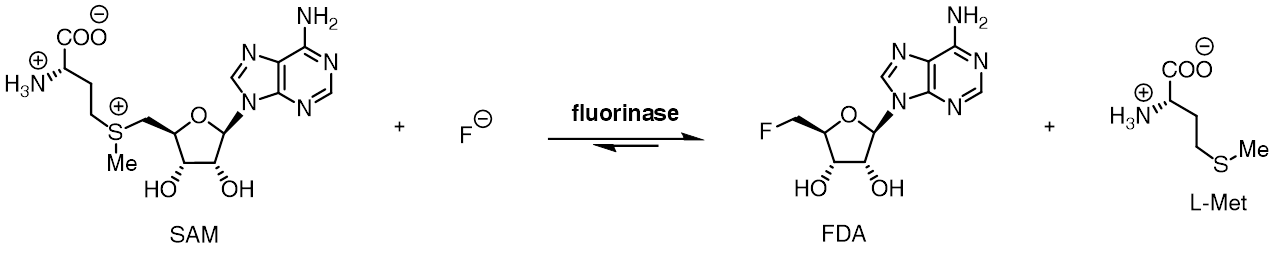
The fluorinase enzyme (, also known as adenosyl-fluoride synthase) catalyzes the reaction between fluoride ion and the co-factor S-adenosyl-L-methionine (SAM) to generate L-methionine and 5'-fluoro-5'-deoxyadenosine, the first committed product of the fluorometabolite biosynthesis pathway. The fluorinase was originally isolated from the soil bacterium Streptomyces cattleya, but homologues have since been identified in a number of other bacterial species, including Streptomyces sp. MA37, Nocardia brasiliensis and Actinoplanes sp. N902-109. This is the only known enzyme capable of catalysing the formation of a carbon-fluorine bond, the strongest single bond in organic chemistry.

Reactivity
The fluorinase catalyses an SN2-type nucleophilic substitution at the C-5' position of SAM, while L-methionine acts as a neutral leaving group. The fluorinase-catalysed reaction is estimated to be between 106 and 1015 times faster than the uncatalysed reaction, a significant rate enhancement. Despite this, the fluorinase is still regarded as a slow enzyme, with a turnover number (kcat) of 0.06 min−1. The high kinetic barrier to reaction is attributed to the strong solvation of fluoride ion in water, resulting in a high activation energy associated with stripping solvating water molecules from aqueous fluoride ion, converting fluoride into a potent nucleophile within the active site.
The reaction catalysed by the fluorinase is reversible, and upon incubation of 5'-fluoro-5'-deoxyadenosine and L-methionine with the fluorinase, SAM and fluoride ion are produced. Replacing L-methionine with L-selenomethionine results in a 6-fold rate enhancement of the reverse reaction, due to the increased nucleophilicity of the selenium centre compared to the sulfur centre.
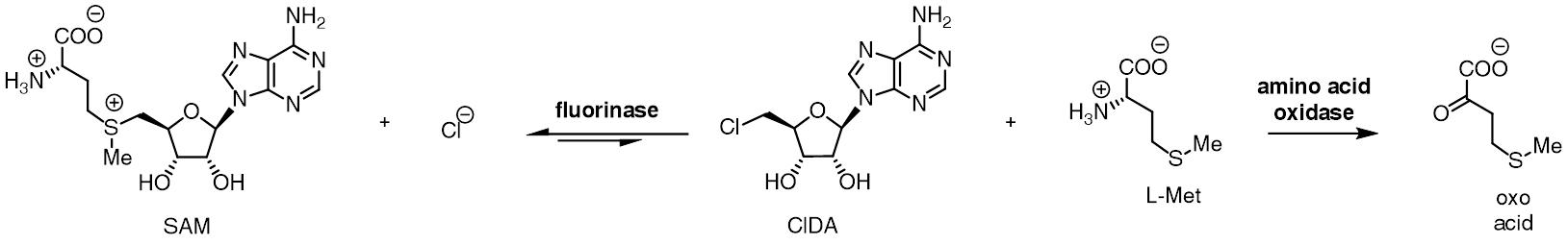
The fluorinase shows a degree of substrate tolerance for halide ion, and can also use chloride ion in place of fluoride ion. While the equilibrium for reaction between SAM and fluoride ion lies towards products FDA and L-methionine, the equilibrium position is reversed in the case for chloride ion. Incubation of SAM and chloride ion with the fluorinase does not result in generation of 5'-chloro-5'-deoxyadenosine (ClDA), unless an additional enzyme, an L-amino acid oxidase, is added. The amino acid oxidase removes the L-methionine from the reaction, converting it to the corresponding oxo-acid.

Structural studies
As of late 2007, 9 structures have been solved for this class of enzymes, with PDB accession codes , , , , , , , , and .
The names given to the enzyme come not from the structure, but from the function: 5-Fluoro-5-deoxyadenosine is the molecule synthesised. The structure is homologous to the duf-62 enzyme series. The enzyme is a dimer of trimers (2 molecules each with three subunits). The active sites are located between these subunits (subunit interfaces), each can bind to one SAM molecule at a time.
Fluorometabolite biosynthesis
References
References
- (February 2014). "Identification of fluorinases from Streptomyces sp MA37, Norcardia brasiliensis, and Actinoplanes sp N902-109 by genome mining". ChemBioChem.
- (February 2008). "Understanding organofluorine chemistry. An introduction to the C-F bond". Chemical Society Reviews.
- (January 2008). "Discovery and characterization of a marine bacterial SAM-dependent chlorinase". Nature Chemical Biology.
- (May 2004). "Enzymatic fluorination in Streptomyces cattleya takes place with an inversion of configuration consistent with an SN2 reaction mechanism". ChemBioChem.
- (October 2005). "Insight into enzymatic C-F bond formation from QM and QM/MM calculations". Journal of the American Chemical Society.
- (October 2013). "Catalysis by desolvation: the catalytic prowess of SAM-dependent halide-alkylating enzymes". Journal of the American Chemical Society.
- (November 2007). "Mechanism of enzymatic fluorination in Streptomyces cattleya". Journal of the American Chemical Society.
- (January 2006). "The fluorinase from Streptomyces cattleya is also a chlorinase". Angewandte Chemie.
- (February 2006). "Fluorinase mediated C-(18)F bond formation, an enzymatic tool for PET labelling". Chemical Communications.
- (September 2015). "A two-step fluorinase enzyme mediated (18)F labelling of an RGD peptide for positron emission tomography". Chemical Communications.
- (February 2004). "Crystal structure and mechanism of a bacterial fluorinating enzyme". Nature.
This article was imported from Wikipedia and is available under the Creative Commons Attribution-ShareAlike 4.0 License. Content has been adapted to SurfDoc format. Original contributors can be found on the article history page.
Ask Mako anything about Fluorinase — get instant answers, deeper analysis, and related topics.
Research with MakoFree with your Surf account
Create a free account to save articles, ask Mako questions, and organize your research.
Sign up freeThis content may have been generated or modified by AI. CloudSurf Software LLC is not responsible for the accuracy, completeness, or reliability of AI-generated content. Always verify important information from primary sources.
Report