From Surf Wiki (app.surf) — the open knowledge base
FAM200A
Protein-coding gene in the species Homo sapiens
Protein-coding gene in the species Homo sapiens
This article is written in AMERICAN ENGLISH. Please DO NOT change the spelling to match your personal preference. If you think there is a compelling reason to change the spelling style, please first read Wikipedia:Manual of Style (spelling) and then begin a discussion on the talk page.--
C7orf38 is a gene located on chromosome 7 in the human genome. The gene is expressed in nearly all tissue types at very low levels. Evolutionarily, it can be found throughout the kingdom animalia. While the function of the protein is not fully understood by the scientific community, bioinformatic tools have shown that the protein bares much similarity to zinc finger or transposase proteins. Many of its orthologs, paralogs, and neighboring genes have been shown to possess zinc finger domains. The protein contains a hAT dimerization domain nears its C-terminus. This domain is highly conserved in transposase enzymes.
Gene
C7orf38 is located on chromosome 7 at q22.1. Its genomic sequence contains 5,612 bp. The predominant transcript contains two exons and is 2,507 bp in length. The translated protein contains 573 amino acids.

Protein composition
The 573 amino acid protein has a molecular weight of 66,280.05. The isoelectric point was found to occur at a pH of 5.775, about 1.6 pH lower than that of the average human pH. Two deviations from prototypical human proteins are evident. The protein contains a less than expected number of glycine residues, and is rich in leucine residues. There are not sections of strong hydrophobicity or hydrophilicity. Thus, it is not predicted to be a transmembrane protein.

Gene neighborhood
The four genes in closest proximity to C7orf38 on chromosome 7 exhibit similar function, many of which are transcription factors.

| Name | Orientation | Function |
|---|---|---|
| ZNF789 | Start: 98,908,451 bp from pter | The gene encodes the zinc finger protein 789. Functionally, the gene has been proposed to participate in regulation of transcription. It is expected to use zinc ion binding. |
| ZNF394 | Start: 98,928,790 bp from pter | The gene encodes zinc finger protein 394. Over expression over ZNF394 inhibits the transchription of c-jun and Ap-1. Suggesting that it is a transcriptional repressor. |
| ZKSCAN5 | Start: 98,940,209 bp from pter | The gene encodes zinc finger with KRAB and SCAN domains 5. This gene encodes a zinc finger protein of the Kruppel family. The protein contains a SCAN box and a KRAB A domain. |
| ZNF655 | Start: 98,993,981 bp from pter | The gene encodes zinc finger protein 655. Numerous alternatively spliced transcripts encoding distinct isoforms have been discovered. |
| Mihuya | Start: 99,149,738 bp from pter | The Mihuya gene does not encode a large or known functional protein. The antisense relationship to C7orf38 raises the possibility for regulation of expression. |
Paralogs
Eight paralogs are found in the human proteome. Similar to the neighboring genes, many of the paralogs function as zinc fingers, or transcription factors.
| Name | NCBI Accession Number | Length (AA) | % Identity to C7orf38 | % Similarity to C7orf38 |
|---|---|---|---|---|
| hypothetical protein LOC285550 | NP_001138663.1 | 657 | 79 | 91 |
| zinc finger MYM-type protein 6 | NP_009098.3 | 1325 | 38 | 60 |
| SCAN domain-containing protein 3 | NP_443155.1 | 1325 | 39 | 60 |
| zinc finger BED domain-containing protein 5 | NP_067034.2 | 692 | 35 | 57 |
| transposon-derived Buster3 transposase-like | NP_071373.2 | 594 | 32 | 53 |
| general transcription factor II-I repeat domain-containing protein 2B | NP_001003795.1 | 949 | 25 | 46 |
| GTF2I repeat domain containing 2 | NP_775808.2 | 949 | 24 | 45 |
| EPM2A interacting protein 1 | NP_055620.1 | 607 | 22 | 42 |
Orthologs
Orthologs to C7orf38 can be traced back evolutionarily through plants. The following is not an extensive list of orthologs. It is intended to provide an evolutionary overview of the conservation of C7orf38.
| Common name | Genus & species | NCBI accession number | Length (AA) | % Identity to C7orf38 | % Similarity to C7orf38 |
|---|---|---|---|---|---|
| Chimp | Pan troglodytes | XP_001139775.1 | 573 | 99 | 99 |
| Macaque monkey | Macaca fascicularis | BAE01234.1 | 573 | 96 | 98 |
| Horse | Equus caballus | XP_001915370.1 | 573 | 81 | 84 |
| Pig | Sus scrofa | XP_001929194 | 1323 | 39 | 61 |
| Cow | Bos taurus | XP_875656.2 | 1320 | 38 | 61 |
| Mouse | Mus musculus | CAM15594.1 | 1157 | 37 | 60 |
| Domestic dog | Canis lupus familiaris | ABF22701.1 | 609 | 37 | 60 |
| Rat | Rattus rattus | NP_001102151.1 | 1249 | 37 | 59 |
| Opossum | Monodelphis domestica | XP_001372983.1 | 608 | 37 | 59 |
| Chicken | Gallus gallus | XP_424913.2 | 641 | 37 | 58 |
| Frog | Xenopus (Silurana) tropicalis | ABF20551.1 | 656 | 37 | 56 |
| Zebra fish | Danio rerio | XP_001340213.1 | 609 | 37 | 56 |
| Pea aphid | Acyrthosiphon pisum | XP_001943527.1 | 659 | 36 | 54 |
| Beatle | Tribolium castaneum | ABF20545.1 | 599 | 35 | 55 |
| Sea squirt | Ciona intestinalis | XP_002119512.1 | 524 | 34 | 52 |
| Hydra | Hydra magnipapillata | XP_002165429.1 | 572 | 29 | 52 |
| Puffer fish | Tetraodon nigroviridis | CAF95678.1 | 539 | 28 | 47 |
| Mosquito | Anopheles gambiae | XP_558399.5 | 591 | 28 | 47 |
| Sea urchin | Strongylocentrotus purpuratus | ABF20546.1 | 625 | 27 | 47 |
| Grass plant | Sorghum bicolor | XP_002439156.1 | 524 | 25 | 40 |
| Broad leaf tree | Populus trichocarpa | XP_002319808.1 | 788 | 21 | 39 |
Structure
Protein
CBLast was used to determine a structurally related protein with experimentally determined structure. The protein Hermes DNA transposase, of the Hermes DBD superfamily, was shown to be structurally similar (Evalue: 1E-6). The hAT dimerization domain is found at the C-terminus of transposase elements belonging to the Activator superfamily (hAT element superfamily). The isolated dimerization domain forms extremely stable dimers in vitro.

mRNA
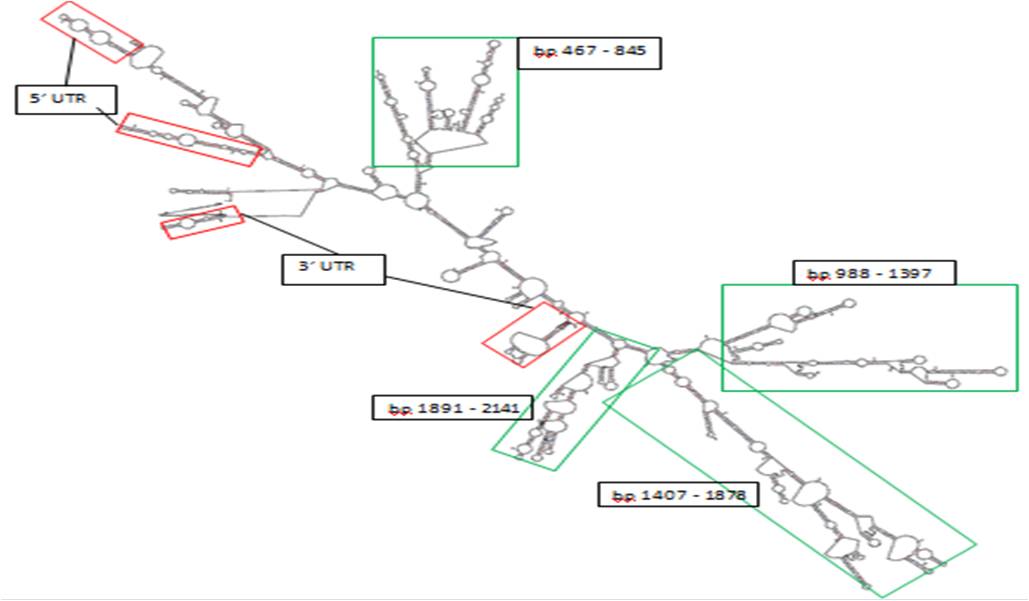
The MFOLD program available at Rensselaer BioInformatics Server was used to predict secondary structure of the mature mRNA sequence. The primary sequence of the mRNA secondary structures displayed high levels of conservation in orthologs, suggesting structural importance.
Tissue distribution
The gene appears to be expressed in most tissue types. Very low levels of expression were observed through est profiles, and no deviation was observed between health or developmental states.



References
References
- "University of California Santa Cruz".
- "NCBI UniGene".
- "NCBI BLAST".
- "KEGG".
- (2000). "A highly conserved domain of the maize activator transposase is involved in dimerization.". Plant Cell.
- (24 February 2010). "Fam200A".
- "NCBI Protein Accession Number".
- (December 2016). "AAStats. SDSC Biology WorkBench".
- (December 2016). "IP. SDSC Biology WorkBench".
- (December 2016). "SAPS. SDSC Biology WorkBench".
- "AceView".
- "Hermes DNA Transposase".
- "Fam200A".
- "NCBI UniGene".
This article was imported from Wikipedia and is available under the Creative Commons Attribution-ShareAlike 4.0 License. Content has been adapted to SurfDoc format. Original contributors can be found on the article history page.
Ask Mako anything about FAM200A — get instant answers, deeper analysis, and related topics.
Research with MakoFree with your Surf account
Create a free account to save articles, ask Mako questions, and organize your research.
Sign up freeThis content may have been generated or modified by AI. CloudSurf Software LLC is not responsible for the accuracy, completeness, or reliability of AI-generated content. Always verify important information from primary sources.
Report